Cell Infection Dynamics

Team Leader
To survive and replicate within a cell, intracellular bacteria, viruses, or parasites, must accommodate the intracellular environment to fulfill their needs. In response to this invasion, the host cell initiates defense mechanisms to eliminate the pathogen. Our research focuses on these highly dynamic interactions between intracellular pathogens and their host cells.
Intracellular pathogens have evolved into inadvertent teachers of cell biology. Through their unique perturbation of normal cell biology, they show us novel aspect of molecular mechanisms controlling dynamic cellular events.

Our emerging research team aims to understand the molecular dynamics that unfold during intracellular infection. Our primary pathogen of interest is Salmonella, a food-borne bacterium classified as “high-priority” by the World Health Organization and a versatile model to study intracellular bacterial pathogenesis.
We investigate how Salmonella hijacks host signaling pathways to remodel intracellular architecture and establish a unique replication-permissive niche that supplies nutrients while limiting host detection. A central focus of our work is how post-translational modifications reshape intracellular trafficking during infection, ultimately driving the formation of a specialized bacteria-containing vacuole that fulfills bacterial needs.
Because intravacuolar replication is a strategy shared by major human pathogens, including Mycobacterium tuberculosis, Chlamydia trachomatis, and Legionella pneumophila, our research provides a framework to identify common principles of niche formation. By uncovering these shared mechanisms, we aim to inform new therapeutic strategies that target the intracellular shelters on which pathogens depend for survival.
Team
Scientists
Virginie Stévenin
Master Student
Esteban Francois
Stévenin et al. (2024) Multi-omic analyses of cancer-associated clinical Salmonella reveal a bacterial-induced host metabolic shift leading to cell transformation. Cell Rep
Stévenin & Neefjes (2022) Control of host PTMs by intracellular bacteria: an opportunity towards novel anti-infective agents. Cell Chem Biol
Stévenin et al. (2021) Purification of infection-associated macropinosomes by magnetic isolation for proteomic characterization. Nat Protoc
Stévenin et al. (2019) Dynamic growth and shrinkage of the Salmonella-containing vacuole determines the intracellular pathogen niche. Cell Rep
Voznica et al. (2018). Identifying parameters of host cell vulnerability during Salmonella infection by quantitative image analysis and modeling. Infect Immun
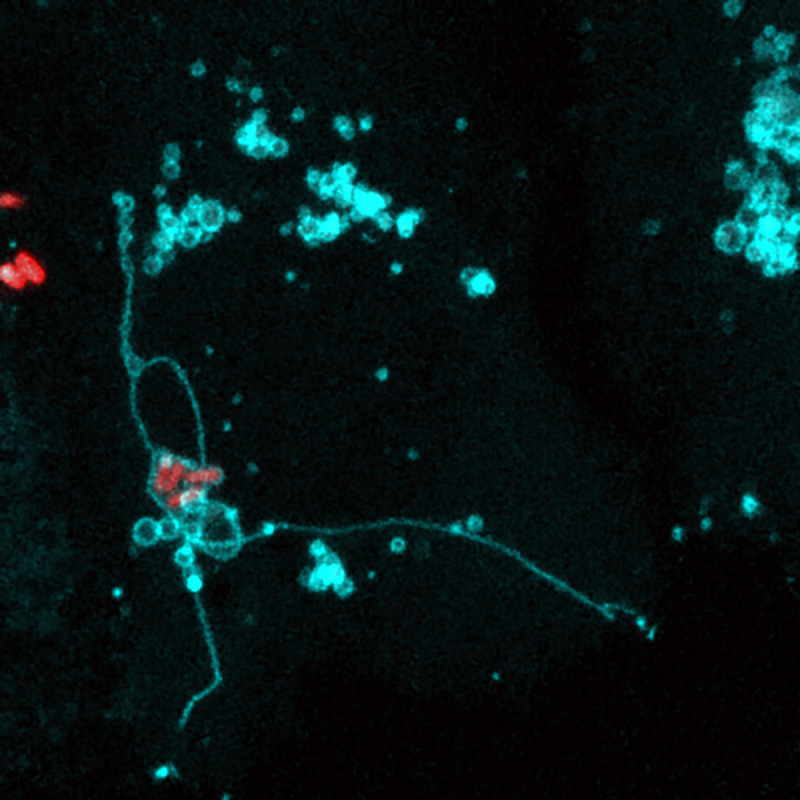
Salmonella building a vacuolar niche in an epithelial cell.
2026
Paradis M#, Gorse L#, Silva Da Costa L, Chua R, Alberola G, Metais A, Batistic L, Benoist T, David A, Pages E, Gomes A, Bomfim C, Ravon-Katossky L, Suire B, Fromont L, Siaden-Ortega R, Pericat D, Cougoule C, Boyer L, Gaudenzio N, Mazars R, Stévenin V, Zhong F, Olagnier D, Meunier E*. (2026) A TAK1-driven NLRP1 inflammasome pathway revealed by phosphatase-targeting environmental toxins. BioRxiv doi:10.64898/2026.01.23.701233
Sapmaz A, van der Zanden S, Akkermans J, Menager L, Janssen L, Francois E, Hagoort N, Berlin I, Neefjes J, Stévenin V*. (2026) Salmonella exploits USP32 to coordinate Rab14 and Rab11 recycling pathways for intracellular survival. BioRxiv doi:10.64898/2026.01.01.696821
2025
De Meyer M, Verhee A, Grzesik H, De Sutter D, Huyghe J, Delhaye L, Van de Steene T, Fijalkowski I, Jonckheere V, Meuris L, Bertrand M, Van Damme P#*, Stévenin V#*, Eyckerman S#*. (2025) Virotrap Reveals Salmonella SopB as A Ubiquitinated Cargo for Host ESCRT-0. BioRxiv doi:10.1101/2025.08.19.669813
Stévenin V, Neefjes J*. (2025) Salmonella, the insidious contributor to gallbladder and colon cancers. Nature Reviews Cancer. doi:10.1038/s41568-025-00812-z
2024
Stévenin V#*, Coipan C#, Duijster J, van Elsland D, Voogd L, Bigey L, van Hoek A, Wijnands L, Jansen L, Akkermans J, Neefjes-Borst A, Franz E, Mughini-Gras L, Neefjes J*. (2024) Multi-omics analyses of cancer-linked clinical salmonellae reveal bacterial-induced host metabolic shift and mTOR-dependent cell transformation. Cell Reports. 43(11):114931. doi:10.1016/j.celrep.2024.114931
2023
Stévenin V*, Neefjes J. (2023). Soft agar colony formation assay to quantify mouse embryonic fibroblast transformation after Salmonella infection. STAR Protocols. 4(3):102379. doi: 10.1016/j.xpro.2023.10237
Berlin I*, Sapmaz A, Stévenin V, Neefjes J. (2023) Ubiquitin and its relatives as wizards of the endolysosomal system. Journal of Cell Science. 136 (4): jcs260101. doi: 10.1242/jcs.260101
2022
van Elsland D#, Duijster J#, Zhang J#, Stévenin V, Zhang Y, Zha L, Xia Y, Franz E, Sun J*, Mughini-Gras L*, Neefjes J*. (2022) Repetitive non-typhoidal Salmonella exposure is an environmental risk factor for colon cancer and tumor growth. Cell Reports Medicine. 3: 100852. doi: 10.1016/j.xcrm.2022.100852
Stévenin V*, Neefjes J. (2022). Control of host PTMs by intracellular bacteria: an opportunity towards novel anti-infective agents. Cell Chemical Biology. 29(5):741-756. doi: 10.1016/j.chembiol.2022.04.004.
Schink K*, Tan KW, Spangenberg H, Martorana D, Sneeggen M, Stévenin V, Enninga J, Campsteijn C, Raiborg C, Stenmark H*. (2021) The phosphoinositide coincidence detector Phafin2 promotes macropinocytosis by coordinating actin organisation at forming macropinosomes. Nature Communications 12(1):6577. doi: 10.1038/s41467-021-26775-x
2021
Stévenin V*, Giai Gianetto Q, Duchateau M, Matondo M, Enninga J, Chang YY*. (2021) Purification of infection-associated macropinosomes by magnetic isolation for proteomic characterization. Nature Protocols 16(11):5220-5249. doi: 10.1038/s41596-021-00610-5
Chang YY*, Enninga J, Stévenin V*. (2021) New methods to decrypt emerging macropinosome functions during the host-pathogen crosstalk. Cellular Microbiology 23(7):e13342. doi: 10.1111/cmi.13342.
2020
Chang YY, Stévenin V, Duchateau M, Giai Gianetto Q, Hourdel V, Rodrigues CD, Matondo M, Reiling N, Enninga J*. (2020) Shigella hijacks the exocyst to cluster macropinosomes for efficient vacuolar escape. PLoS Pathogens. 16(8):e1008822. doi: 10.1371/journal.ppat.1008822.
2019
Stévenin V, Enninga J*. (2019) Cellular imaging of intracellular bacterial pathogens. Microbiology Spectrum 7(2). doi:10.1128/microbiolspec.BAI-0017-2019
Stévenin V, Chang YY, Le Toquin Y, Duchateau M, Gianetto QG, Luk CH, Salles A, Sohst V, Matondo M, Reiling N, Enninga J*. (2019) Dynamic growth and shrinkage of the Salmonella-containing vacuole determines the intracellular pathogen niche. Cell Reports. 29(12):3958-3973.e7. doi: 10.1016/j.celrep.2019.11.049.
2018
Voznica J, Enninga J, Stévenin V*. (2018) High-throughput microscopic analysis of Salmonella invasion of host cells. Bio-protocol 20(18):e3017. doi: 10.21769/BioProtoc.3017.
Fredlund J#, Santos JC#, Stévenin V#, Weiner A, Rechav K, Mallet A, Krijnse-Locker J, Elbaum M, Enninga J*. (2018) The early entry of Salmonella in a distinct tight compartment revealed at high temporal and ultrastructural resolution. Cellular Microbiology. 20(4):e12816. doi: 10.1111/cmi.12816.
Voznica J, Gardella C, Belotserkovsky I, Dufour A, Enninga J, Stévenin V*. (2018) Identifying parameters of host cell vulnerability during Salmonella infection by quantitative image analysis and modeling. Infection and Immunity. 86(1):e00644-17. doi: 10.1128/IAI.00644-17
Collaborations
Etienne Meunier, IPBS, Toulouse, France
Agnès Wiedemann, IRSD, Toulouse, France
Pedro Escoll, Institut Pasteur, Paris, France
Jost Enninga, Institut Pasteur, Paris, France
Kristine Schauer, Institut Gustave Roussy, Paris, France
Etienne Morel, Institut Necker Enfants Malades, Paris, France
Ting-Di Wu, Institut Curie, Paris, France
Franck Perez, Institut Curie, Paris, France
Sven Eyckerman, VIB-Ghent, Belgium
Petra van Damme, VIB-Ghent, Belgium
Adán Pinto-Fernández, CAMS Oxford Institute, UK
Jacques Neefjes, LUMC, Leiden, the Netherlands
Mikael Sellin, Uppsala University, Sweden
Funding
- VENI grant from the NWO Talent Programme (2025-2027)
- « Amorçage de jeunes équipes » program from the Fondation for Medical Research (FRM, 2026-2029)


