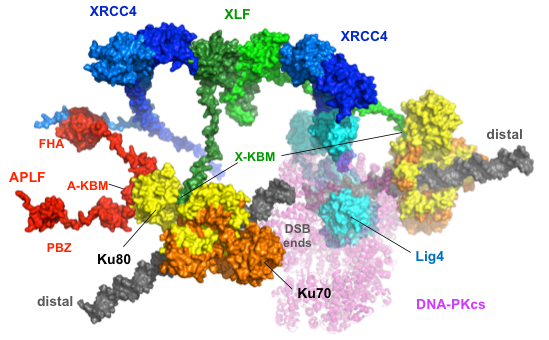
Characterization of non-homologous end joining
NHEJ is initiated by recognition of the DSBs ends by the Ku70-Ku80 heterodimer. Ku/DNA ends then serve as a hub for the recruitment of a DNA-dependent serine-threonine kinase (DNA-PK) and of other factors involved in ends processing that prepares their final ligation by the XLF-XRCC4-ADN Ligase 4 complex (Frit et al. 2019). In line with this research topic, our main recent discoveries are:
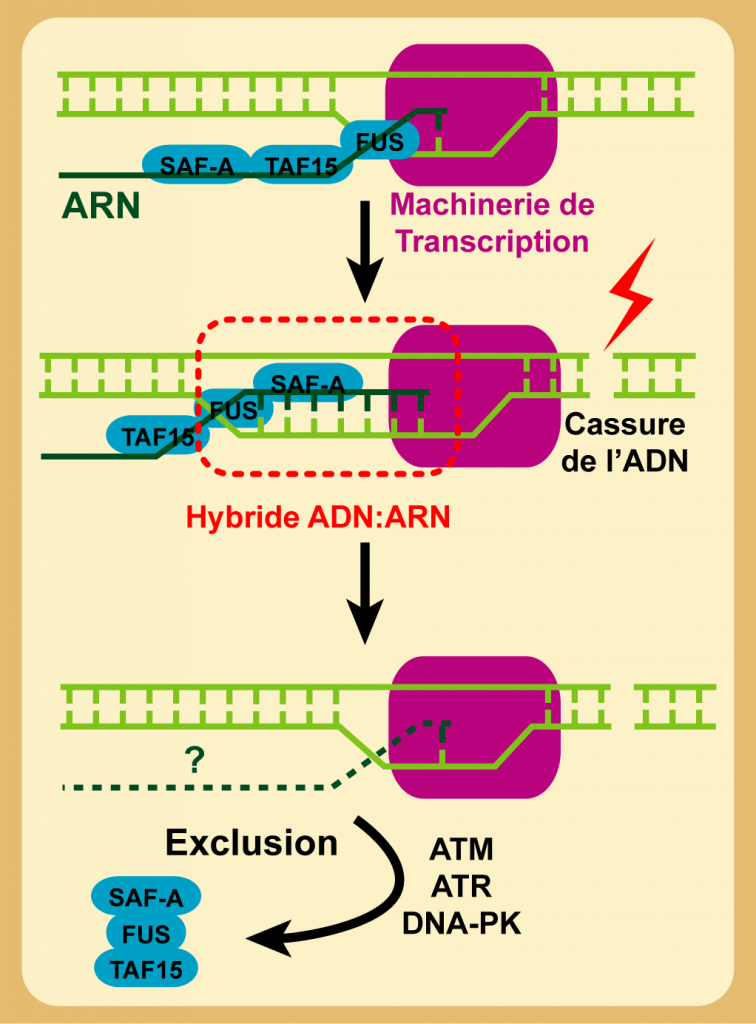
R-loops regulation at DSBs sites – We characterized previously SAF-A, a protein involved in mRNA metabolism, as a novel DNA-PK substrate (Britton et al. 2009). Then, by studying cell mobilization in response to DSBs of SAF-A partners isolated by mass spectrometry, we first characterized a complex behaviour at sites of DSBs with of PARP-dependent recruitment of these proteins superimposed to their exclusion from the damaged areas. We showed that the latter phase is due to an original mechanism resolving RNA:DNA (R-loops) hybrids secondary to damage in transcript DNA (Britton et al. 2014).
Building the NHEJ macromolecular complex – In collaboration with a team specialized in structural biology (JB Charbonnier, Saclay, France), we recently explored some protein-protein interactions within the NHEJ complex. We have defined the precise interaction sites between two NHEJ proteins (APLF and XLF) and the DNA-end binding factor Ku. We characterized that XRCC4, partner of APLF, and XLF present independent recruitment on DSBs, by the respective binding of their A- and X-Ku binding motifs (KBM) to remote sites in Ku80. By using cell lines expressing mutant proteins, we validated in cellulo the key positions on the interaction sites deduced from the resolved corresponding structures. Finally, by using transfected or integrated C-NHEJ reporter substrates in these different cell lines, we showed that combined Ku80 mutations at the binding sites of X-KBM and A-KBM compromise both the efficacy and accuracy of C-NHEJ repair and consequently increase cellular radiosensitivity (Nemoz et al. 2018).