Immuno-modulation by Mycobacterial Lipids and Glycoconjugates

Group Leader
Our main objective is to decipher the molecular cross-talk that establishes between Mycobacterium tuberculosis and host immune cells during infection. We focus on understanding the role and the structure-function relationships of cell envelope lipids and glycoconjugates in innate immune recognition of mycobacteria and stimulation of CD1-restricted T cells.
We combine multidisciplinary approaches to decipher and exploit the immunomodulatory properties of Mycobacterium tuberculosis lipids and glycoconjugates.

We use a multidisciplinary approach combining lipidomics, microbiology, cell biology and animal experimentation to explore at the molecular level several facets of the host-pathogen interaction. Moreover, we wish to translate this fundamental knowledge into new therapeutic, preventive or diagnostic strategies/tools to be explored.
Modulation of innate immune responses
Mycobacterium tuberculosis (Mtb) inhibits innate immune responses, including inflammation and autophagy, thus favoring its survival inside the infected host. Our goals are to uncover the molecular bases of Mtb recognition by pattern-recognition receptors, including TLRs and C-type lectins, and to identify Mtb factors that modulate macrophage function. Moreover, we wish to characterize Mt lipid trafficking within extracellular vesicles and the remodeling of Mtb lipidome during infection.
Glycolipid antigen presentation to CD1-restricted T cells
Lipids and glycolipids are important antigens that induce CD1-restricted T cell-mediated specific immune responses. We intend to define the repertoire of Mtb lipid T cell epitopes, to characterize the molecular steps and mechanisms of their presentation by antigen-presenting cells and to evaluate of the protective efficacy of these antigenic lipids in animal models of Mtb infection.
Structural and functional definitions of Mtb glycoproteome
We previously evidenced that protein-O-mannosylation is crucial for virulence of Mtb. Our aims are to identify and decipher the roles of the mannoproteins contributing to Mtb persistence, in vivo, and to understand the structure/function relationships of protein-O-mannosyl transferase in order to guide inhibition strategies.
Team members
Research Scientists
Emeline Fabre (University)
Martine Gilleron (CNRS)
Emilie Layre (CNRS)
Jérôme Nigou (CNRS)
Virginie Puech (University)
Michel Rivière (CNRS)
Alain Vercellone (University)
Isabelle Vergne (CNRS)
Albertus Viljoen (CNRS)
Research Engineers
Yasmine Achour
Marion Retailleau
Sophie Zuberogoitia (CNRS)
Postdoctoral Fellow
Maxime Megret
PhD Students
Héctor Mayoral Reyes
Chloé Rivière
Viljoen et al. (2023) Nanoscale clustering of mycobacterial ligands and DC-SIGN host receptors are key determinants for pathogen recognition. Sci Adv
Mosquera-Restrepo et al. (2022) A Mycobacterium tuberculosis fingerprint in human breath allows tuberculosis detection. Nat Commun
Blanc et al. (2017) Mycobacterium tuberculosis inhibits human innate immune responses via the production of TLR2 antagonist glycolipids. Proc Natl Acad Sci USA
Decout et al. (2017) Rational design of adjuvants targeting the C-type lectin Mincle. Proc Natl Acad Sci USA
Gilleron et al. (2016) Lysosomal lipases PLRP2 and LPLA2 process mycobacterial multiacylated lipid antigens and generate T cell stimulatory antigens. Cell Chem Biol
Liu, Tonini et al. (2013) Bacterial protein-O-mannosylating enzyme is crucial for virulence of Mycobacterium tuberculosis. Proc Natl Acad Sci USA
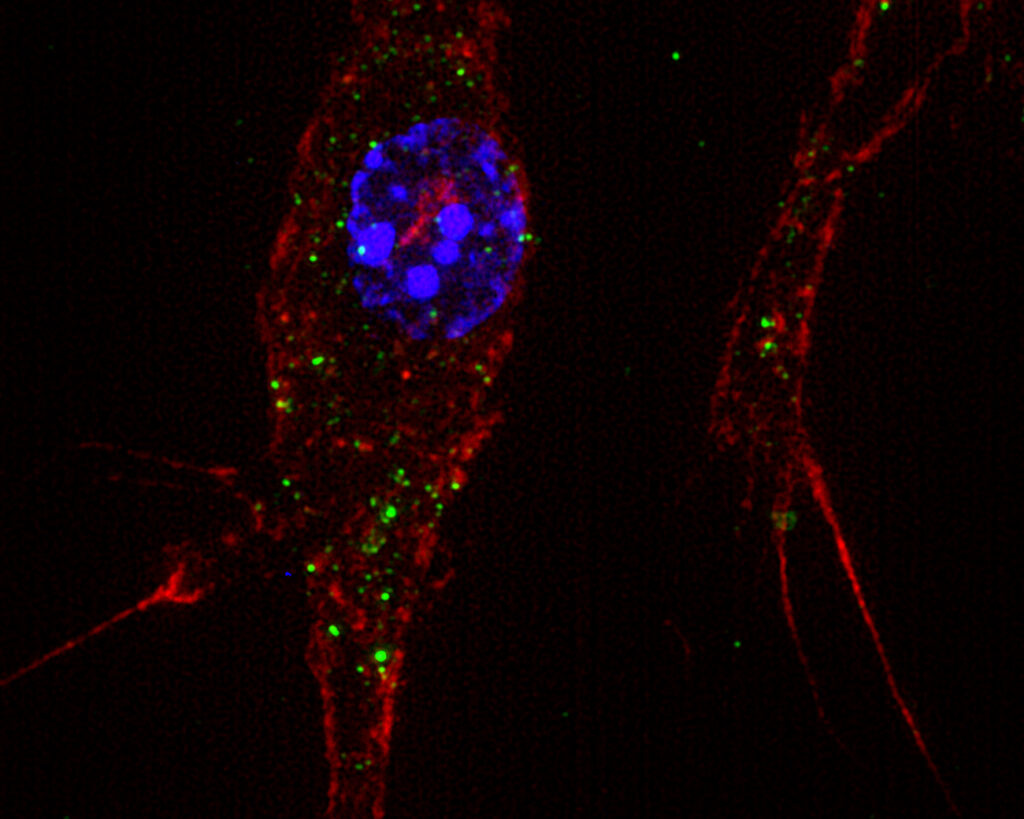
Extracellular vesicles shuttle mycobacterial immunomodulatory lipids toward bystander macrophages. CFSE-labeled M. bovis BCG extracellular vesicles (green), labeled BMDM: nucleus (blue), actin network (red). © Pierre Boyer, Elodie Vega, Antonio Peixoto, Emilie Layre, Jérôme Nigou│IPBS│CNRS/UPS
May 2023: How the organization of carbohydrate molecules on the surface of its envelope allows Mycobacterium tuberculosis to trick the immune system
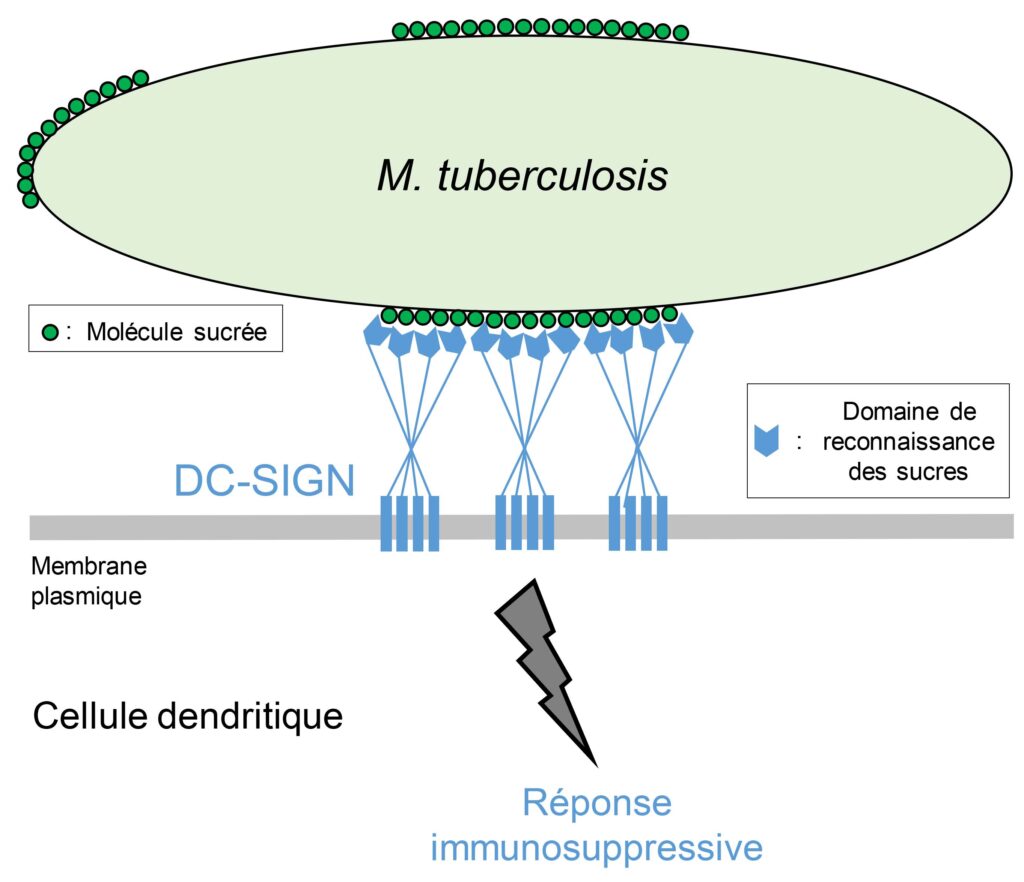
The surface of the envelope of Mycobacterium tuberculosis is covered with carbohydrate molecules that are organized in nano-domains constituting platforms of strong interaction with the DC-SIGN receptor, itself organized in clusters at the membrane of immune cells. The recognition by DC-SIGN induces an immunosuppressive response that promotes the multiplication of the pathogen within the infected organism. Copyright: Jérôme Nigou Thumbnail Copyright: Albertus Viljoen et Yves Dufrêne
In this study, the scientists showed, notably through the use of a molecular imaging technique with nanometric resolution, that the carbohydrate molecules on the surface of the M. tuberculosis complex species, contrary to species of the same bacterial genus, are organized in large domains that constitute platforms of strong interaction with the DC-SIGN receptor, itself organized in clusters.
These results constitute a conceptual advance in the understanding of the functioning of receptors specialized in the recognition of carbohydrate molecules, demonstrating that the organization of ligands at the surface of microorganisms is a key parameter. Moreover, they change our perception of the surface of bacterial envelopes, by suggesting the presence of functional domains. Finally, they open the way to the development of new anti-tuberculosis approaches using immunomodulation.
Source
Nanoscale clustering of mycobacterial ligands and DC-SIGN host receptors are key determinants for pathogen recognition. Albertus Viljoen†, Alain Vercellone†, Myriam Chimen, Gérald Gaibelet, Serge Mazères, Jérôme Nigou* and Yves F. Dufrêne*. Science Advances, 9(20):eadf9498. DOI: 10.1126/sciadv.adf9498. † co-first authors; * co-senior authors- Access the article
Researcher contact
Jérôme Nigou | Jerome.Nigou@ipbs.fr | +33 (0)5 61 17 55 54 | @NigouJ
Press contact
Francoise Viala | communication@ipbs.fr | +33 (0)6 01 26 52 59
Read the Press release (in French): here
December 2022: Towards a point-of-care diagnosis of tuberculosis, directly from the analysis of patients’ exhaled air
The World Health Organization (WHO) estimates that there are 10 million new cases of tuberculosis per year, 12% of which are children. However, 1/3 of these cases remain undiagnosed, mainly because of limitations of the tools currently available. The scientists’ work shows that the air exhaled by tuberculosis patients contains traces of the presence of the bacteria responsible for tuberculosis and that its analysis would allow a much more sensitive, rapid and easy diagnosis of the disease. The study was published on December 14, 2022 in the journal Nature Communications.
WHO’s goal is to reduce TB mortality and incidence by 90% and 80% respectively by 2030 compared to 2015 levels. This cannot be achieved without the development of much more effective diagnostic tools. TB diagnosis is currently based on clinical and radiographic examinations, coupled with bacteriological analysis of sputum where the presence of Mycobacterium tuberculosis, the bacterium responsible for the disease, is searched for by microscopic analysis, culture and/or gene amplification. However, sputum analysis has a low sensitivity in children and in individuals with low sputum counts, such as HIV co-infected patients.
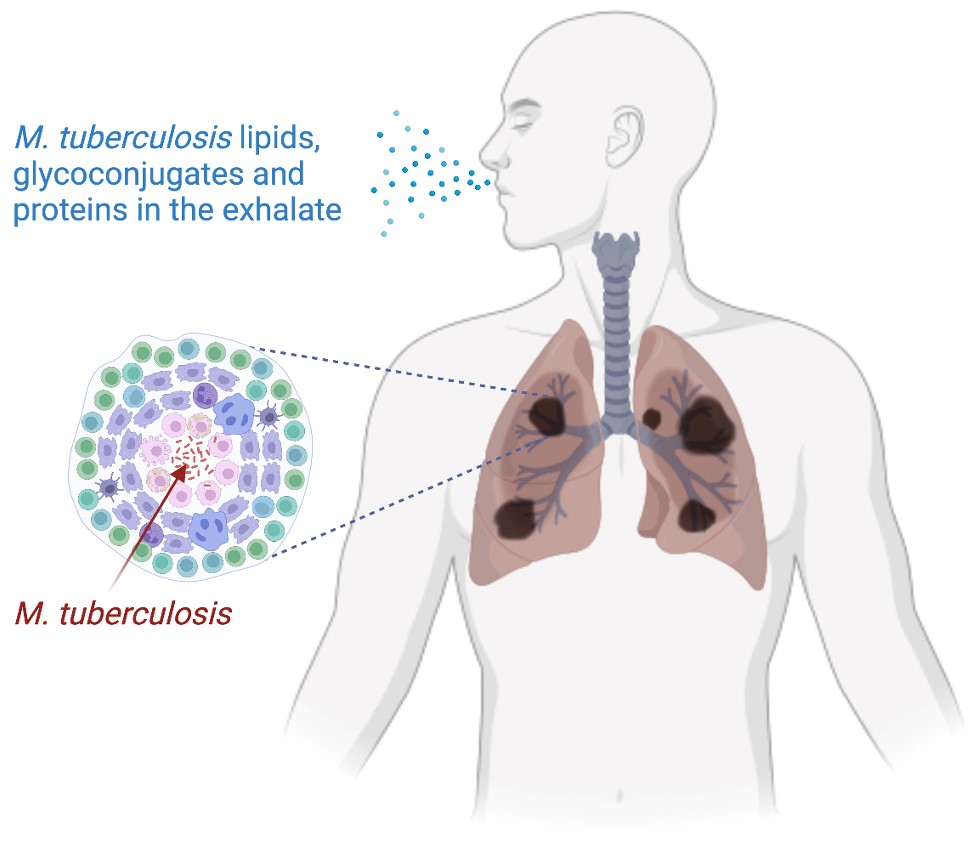
Tuberculosis patients exhale large quantities of Mycobacterium tuberculosis molecules, including lipids, glycoconjugates and proteins, that can be assayed for disease detection. (Created with BioRender.com) © Jérome Nigou, Olivier Neyrolles
In this study, scientists evaluated the diagnostic potential of a poorly explored biological sample, the exhaled breath condensate. This sample, which corresponds to the liquid phase of exhaled air sampled by cooling, has many advantages in the context of TB diagnosis. Indeed, it is constituted by the aerosolization of the liquid covering the respiratory epithelium; its composition thus varies according to the pathological state of the lung. Moreover, its collection is easy, non-invasive, and does not require specialized personnel. Here, scientists have been able to detect in the exhaled breath condensate collected from TB patients the presence of molecules (glycoconjuugates, lipids and proteins) produced by M. tuberculosis, allowing to differentiate these patients from healthy individuals or those with other respiratory infections. Interestingly, this approach, which can be implemented in the short term directly at point-of-care, allows the detection of TB patients for whom the bacteriological analysis of sputum is negative.
The results of this pilot study have already attracted the attention of several national and international organizations and are being validated on larger cohorts of patients in different regions of the world.
Source
A Mycobacterium tuberculosis fingerprint in human breath allows tuberculosis detection” Sergio Fabián Mosquera-Restrepo, Sophie Zuberogoïtia, Lucie Gouxette, Emilie Layre, Martine Gilleron, Alexandre Stella, David Rengel, Odile Burlet-Schiltz, Ana Cecilia Caro, Luis F. Garcia, César Segura, Carlos Alberto Peláez Jaramillo, Mauricio Rojas*, Jérôme Nigou* Nature Communications December 14, 2022 – Access the article
Researcher contact
Jérôme Nigou | Jerome.Nigou@ipbs.fr | +33 (0)5 61 17 55 54 | @NigouJ
Press contact
Francoise Viala | communication@ipbs.fr | +33 (0)6 01 26 52 59
Read the Press release (in French): here
Listen to the podcast on France Culture (in French): here
Read comments in Le Monde: here
October 2017: Mycobacterium tuberculosis plays hide-and-seek with the immune system
To secure their colonization and survival, pathogens have evolved tactics to undermine host immune responses. Better understanding pathogen invasion strategies is key to develop appropriate preventive or therapeutic tools. Scientists from the team “Immunomodulation by Mycobacterial Lipids and Glycoconjugates” at the Institute of Pharmacology and Structural Biology (IPBS – CNRS/Université Toulouse III – Paul Sabatier) have uncovered a molecular mechanism by which the tuberculosis bacillus escapes detection by the innate immune system. This study is published in the Proceedings of the National Academy of Sciences of the USA, on October 2 2017.
Mycobacterium tuberculosis, the causative agent of human tuberculosis, is a bacterial pathogen that has adapted to subvert the function of macrophages, whose one of the roles is to recognize and kill invading microorganisms. In the present study, the scientists used a library of M. tuberculosis mutants to infect macrophages and decipher molecular mechanisms by which the pathogen modulates the function of these immune cells. They found that M. tuberculosis produces cell envelope glycolipids that are antagonists of a macrophage receptor, named TLR2, which is dedicated to the recognition of pathogens, thereby preventing its efficient recognition by the immune system.
Source
Landry Blanc, Martine Gilleron, Jacques Prandi, Ok-ryul Song, Mi-Seon Jang, Brigitte Gicquel, Daniel Drocourt, Olivier Neyrolles, Priscille Brodin, Gérard Tiraby, Alain Vercellone & Jérôme Nigou (2017) Mycobacterium tuberculosis inhibits human innate immune responses via the production of TLR2 antagonist glycolipids. PNAS, Published ahead of print October 2, 2017 doi:10.1073/pnas.1707840114. Access the article
This work was performed in collaboration with the Centre d’Infection et d’Immunité de Lille (CIIL), the Pasteur Institutes from Paris and Korea, and the biotech company InvivoGen (Toulouse).
Researcher contact
Contact Jérôme Nigou (jerome.nigou@ipbs.fr)
Read the Press release (in French): here
February 2017: Rational design of adjuvants targeting the C-type lectin Mincle
Vaccination is a highly effective method of preventing infectious diseases. However, adjuvants are needed to enhance the immunogenicity of vaccine antigens. For many years, the development of adjuvants has been an empirical process. Scientists from the team “Immunomodulation by Mycobacterial Lipids and Glycoconjugates” at the Institute of Pharmacology and Structural Biology show that new generation adjuvants can be designed on a rational basis. This study is published in the Proceedings of the National Academy of Sciences of the USA, on 21 Feb 2017.
Vaccines are mainly used to fight infectious diseases but they can also help to treat cancer. Historically, vaccines have been made with live attenuated or dead pathogens. For many pathogens, however, such vaccines have not been successfully developed. Moreover, they can be associated to side effects. Current efforts aim at the development of subunit vaccines that induce a well-defined immune response against specific antigens, from pathogens (virus, bacteria) as well as from tumors.
However, vaccine antigens are often poorly immunogenic and require additional components, termed adjuvants, to help stimulate protective immunity. Most of the adjuvants in use have been developed empirically without a clear understanding of their mechanisms of action. Adjuvants inducing a strong humoral immunity, which protects against extracellular pathogens, are available. But, adjuvants directing the development of robust cellular immune responses, required to fight against intracellular pathogens or tumors, are still needed. Recently, the C-type lectin receptor Mincle was found to elicit such responses on the recognition of microbial glycolipids, thereby providing a basis for the rational design of new adjuvants. In the present study, the scientists used a multidisciplinary approach, combining chemical synthesis, cell biology, and molecular modeling to decipher the molecular bases of ligand recognition by the receptor. This led them to synthesize new compounds inducing stronger immune responses, while being less toxic, than the currently available Mincle ligands, and that represent new powerful adjuvant molecules.
This work was performed in collaboration with the biotech company Invivogen (Toulouse), the “Laboratoire d’Ingénierie des Systèmes Biologiques et des Procédés” (Toulouse) and the “Centro Nacional de Investigaciones Cardiovasculares Carlos III (Madrid).
Source
Decout, A., Silva-Gomes, S., Drocourt, D., Barbe, S., André, I., Cueto, F.J., Lioux, T., Sancho, D., Pérouzel, E., Vercellone, A., Prandi, J., Gilleron, M., Tiraby, G. & Nigou, J. (2017) Rational design of adjuvants targeting the C-type lectin Mincle. PNAS, doi: 10.1073/pnas.1612421114. Access the article
Researcher contact :
Jérôme Nigou | Jerome.Nigou@ipbs.fr | +33 (0)5 61 17 55 54 | @NigouJ
Read the Press release (in French): here
January 2014: A group in Toulouse is awarded a prestigious grant to develop a new vaccine against tuberculosis
Our group has just received a funding from the Brigham & Women’s Hospital, as part of a grant from the Grand Challenges TB Vaccine Accelerator, an initiative of the Bill & Melinda Gates Foundation. This international project involving two american groups, Boston and Atlanta, and three european groups, Salisbury (UK), Groningen (Netherlands) and Toulouse, will explore a very original track to develop a new vaccine against tuberculosis and will be funded to the tune of more than $ 3 million by the foundation. In contrast to classical approaches, it aims at developing a vaccine, which is not based on proteins but on bacterial lipids.
May 2013: Mannodendrimers : anti-inflammatory drugs of tomorrow ?
The discovery of an immune escape strategy used by Mycobacterium tuberculosis, the etiological agent of human tuberculosis, has led the team “Immunochemistry and mycobacterial glycoconjugates” from Institut de Pharmacologie et de Biologie Structurale (IPBS, CNRS/Université Toulouse III – Paul Sabatier) to design a novel type of powerful anti-inflammatory molecules. Published in PNAS on May 13, this work performed in collaboration with the Laboratoire de Chimie de Coordination (LCC, CNRS/UPS/INPT) and Laboratoire de Toxicologie Alimentaire (TOXALIM, INRA/UPS/INPT) opens avenues for the development of new therapeutic strategies to combat inflammatory diseases.
Read the Press release (in French): here
April 2013: Bacterial protein-O-mannosylating enzyme is crucial for virulence of Mycobacterium tuberculosis
A posttranslational protein O-mannosylation process resembling that found in fungi and animals has been reported in the major human pathogen Mycobacterium tuberculosis (Mtb) and related actinobacteria. However, the role and incidence of this process, which is essential in eukaryotes, have never been explored in Mtb.
Read the Press release (in French): here
Collaborations
International
- L. Balboa Instituto de Investigaciones Biomédicas en Retrovirus y Sida, Argentina
- M. Bastian Paul Ehrlich-Institut, Langen, Germany
- M. Behr McGill University Health Centre Research Institute, Montréal, Canada
- V. Dongkeng Centre Pasteur Caméroun, Yaoundé, Cameroun
- M. Dyrhol-Riise University of Oslo, Oslo, Norway
- M.A. Florez-Valdez Centro de Investigación y Asistencia en Tecnología y diseño del Estado de Jalisco, Mexico
- M. Guerin Spanish National Research Council, Barcelona, Spain
- M. Jackson Colorado State University, Fort Collins, USA
- G. Källenius Karolinska Institutet, Stockholm, Sweden
- M. Khaja Mafij Uddin International Centre for Diarrhoeal Disease Research Bangladesh, Dhaka, Bangladesh
- R. Martínez-Máñez Universitat de València, Valencia, Spain
- N. Martinson University of the Witwatersrand, Johannesburg, South Africa
- A. Minnaard University of Groningen, Groningen, The Netherlands
- D.B. Moody Brigham and Women’s Hospital, Boston, USA
- D. D. Pinschewer University of Basel, Basel, Switzerland
- A. Pinter Rutgers University, Newark, USA
- N. Rakotosamimanana Institut Pasteur de Madagascar, Antananarivo, Madagascar
- H. Ramachandraiah FIND, Geneva, Switzerland
- Y. Reiter Technion-Israel Institute of Technology, Haifa, Israel
- M. Rojas University of Antioquia, Medellin, Colombia
- M. Ruhwald FIND, Geneva, Switzerland
- C. Seshadri University of Washington, Seattle, USA
- S. Stenger Institut für Med. Mikrobiologie und Hygiene, Ulm, Germany
- N. Tukvadze National Center for Tuberculosis and Lung Diseases (NCTLD), Tbilisi, Georgia
- S. Vashakidze National Center for Tuberculosis and Lung Diseases (NCTLD), Tbilisi, Georgia
- C. Vilaplana Institut i Hospital d’Investigació en Ciències de la Salut i Hospital Germans Trias i Pujol, Badalona, Spain
- D. Warner University of Cape Town, Cape Town, South Africa
- R. Wood Desmond Tutu Health Foundation NPC, Mowbray, South Africa
National
- I. André CNRS, INSA, Université de Toulouse
- Y. Bourne CNRS, Université de Marseille
- M. Breban INSERM, Université de Versailles Saint-Quentin en Yvelines
- O. Burlet-Schiltz IPBS
- S. Cannaan CNRS, Université de Marseille
- M. Chavent IPBS
- O. Dumitrescu Hospices Civils de Lyon
- G. Ferré IPBS
- C. Guilhot IPBS
- J. Hoffmann and S. Pouzol Fondation Mérieux, Lyon, France
- L. Herrmann INSERM, Université de Versailles Saint-Quentin en Yvelines
- L. Kremer CNRS, Université de Montpellier
- E. Meunier IPBS
- O. Neyrolles IPBS
- E. Oswald and P. Branchu IRSD, Toulouse
- L. Poillet-Perez CRCT, Toulouse
Industry
- Invivogen Toulouse, France
- Innovative Diagnostics Grabels, France
- Lionex Diagnostics and Therapeutics Braunschweig, Germany
- PHC Europe B.V. Breda, The Netherlands
Funding
Our team is or has been supported in the past ten years by grants or fellowships from:
International
- Bill & Melinda Gates Foundation (BMGF)
- Centre Franco-Indien pour la Promotion de la Recherche Avancée (CEFIPRA/IFCPAR)
- Conseil National de la Science et de la Technologie du Mexique (CONACYT)
- Chinese Scholarship Council (CSC)
- European Union (EU)
- Federation of European Biochemical Societies (FEBS)
- National Institute of Health (NIH, USA)
- Swedish Research Council (Vetenskapsrådet)
National
- Agence Nationale de la Recherche (ANR)
- Agence Nationale de la Recherche sur le SIDA et les Maladies Infectieuses Emergentes (ANRS-MIE)
- Association Nationale de la Recherche et de la Technologie (ANRT)
- Club Francophone de l’Autophagie (CFATG)
- Centre National de la Recherche Scientifique (CNRS)
- Direction Générale de l’Armement (DGA)
- Région Occitanie (web)
- Toulouse Tech Transfer (TTT)
- Université Paul Sabatier de Toulouse (UPS)
Charities
- Fondation pour la Recherche Médicale (FRM) Our team was labelled by the Fondation pour la Recherche Médicale (2018-2021)
- Vaincre la Mucoviscidose (VLM)
- Fondation Roland Garrigou (Fonroga)
Industry
Publications
The complete list of our publications is available through Pubmed
Modulation of innate immune response by mycobacteria
Viljoen, A., Vercellone, A., Chimen, M., Gaibelet G., Mazères, S, Nigou, J., Dufrêne, Y.F. (2023) Nanoscale clustering of mycobacterial ligands and DC-SIGN host receptors are key determinants for pathogen recognition. Sci Adv 9, eadf9498 (View)
Mosquera-Restrepo, S.F., Zuberogoitia, S., Gouxette, L., Layre, E., Gilleron, M., Stella, A., Rengel, D., Burlet-Schiltz, O., Caro, A.C., Garcia, L.F., Segura, C., Pelaez Jaramillo, C.A., Rojas, M., Nigou, J. (2022) A Mycobacterium tuberculosis fingerprint in human breath allows tuberculosis detection. Nat Commun 13, 7751 (View)
Bah, A., Sanicas, M., Nigou, J., Guilhot, C., Astarie-Dequeker, C., Vergne, I. (2020) The Lipid Virulence Factors of Mycobacterium tuberculosis Exert Multilayered Control over Autophagy-Related Pathways in Infected Human Macrophages. Cells 9, 666 (View)
Blanc, L., Gilleron, M., Prandi, J., Song, O.R., Jang, M.S., Gicquel, B., Drocourt, D., Neyrolles, O., Brodin, P., Tiraby, G., Vercellone, A., Nigou, J. (2017) Mycobacterium tuberculosis inhibits human innate immune responses via the production of TLR2 antagonist glycolipids. Proc Natl Acad Sci USA 114, 11205-11210 (View)
Decout A., Silva-Gomes S., Drocourt D., Barbe S., André I., Cueto F.J., Lioux T., Sancho D., Pérouzel E., Vercellone A., Prandi J., Gilleron M., Tiraby G. & Nigou J. (2017) Rational design of adjuvants targeting the C-type lectin Mincle. Proc Natl Acad Sci USA 114:2675-2680 (View)
Blattes, E., Vercellone, A., Eutamene, H., Turrin, C.O., Theodorou, V., Majoral, J.P., Caminade, A.M., Prandi, J., Nigou, J., Puzo, G. (2013) Mannodendrimers prevent acute lung inflammation by inhibiting neutrophil recruitment. Proc Natl Acad Sci USA 110, 8795-8800 (View)
Liu, C.F., Tonini, L., Malaga, W., Beau, M., Stella, A., Bouyssie, D., Jackson, M.C., Nigou, J., Puzo, G., Guilhot, C., Burlet-Schiltz, O., Riviere, M. (2013) Bacterial protein-O-mannosylating enzyme is crucial for virulence of Mycobacterium tuberculosis. Proc Natl Acad Sci USA 110, 6560-6565 (View)
Lipid presentation to CD1-restricted T cells
Gilleron, M., Lepore, M., Layre, E., Cala-De Paepe, D., Mebarek, N., Shayman, J.A., Canaan, S., Mori, L., Carriere, F., Puzo, G., De Libero, G. (2016) Lysosomal Lipases PLRP2 and LPLA2 Process Mycobacterial Multi-acylated Lipids and Generate T Cell Stimulatory Antigens. Cell Chem Biol 23, 1147-1156 (View)
Garcia-Alles, L.F., Collmann, A., Versluis, C., Lindner, B., Guiard, J., Maveyraud, L., Huc, E., Im, J.S., Sansano, S., Brando, T., Julien, S., Prandi, J., Gilleron, M., Porcelli, S.A., de la Salle, H., Heck, A.J., Mori, L., Puzo, G., Mourey, L., De Libero, G. (2011) Structural reorganization of the antigen-binding groove of human CD1b for presentation of mycobacterial sulfoglycolipids. Proc Natl Acad Sci USA 108, 17755-17760 (View)
Guiard, J., Collmann, A., Gilleron, M., Mori, L., De Libero, G., Prandi, J., Puzo, G. (2008) Synthesis of diacylated trehalose sulfates: candidates for a tuberculosis vaccine. Angew Chem Int Ed Engl 47, 9734-9738 (View)
De la Salle, H., Mariotti, S., Angenieux, C., Gilleron, M., Garcia-Alles, L.F., Malm, D., Berg, T., Paoletti, S., Maitre, B., Mourey, L., Salamero, J., Cazenave, J.P., Hanau, D., Mori, L., Puzo, G., De Libero, G. (2005) Assistance of microbial glycolipid antigen processing by CD1e. Science 310, 1321-1324 (View)
Gilleron, M., Stenger, S., Mazorra, Z., Wittke, F., Mariotti, S., Bohmer, G., Prandi, J., Mori, L., Puzo, G., De Libero, G. (2004) Diacylated sulfoglycolipids are novel mycobacterial antigens stimulating CD1-restricted T cells during infection with Mycobacterium tuberculosis. J Exp Med 199, 649-659 (View)
Biosynthesis and structural analysis of lipids and glycoconjugates
Géraud N., Rivière C., Falcou C., Cioci G., Froment C., Gervais V., Marcoux J., Gilleron M., Nigou J., Fabre E., Rivière M. (2025) Structural insights into the protein mannosyltransferase from Mycobacterium tuberculosis reveal a WW-domain-like protein motif in bacteria. Commun Biol 8, 1175 Pubmed
Géraud N, Falcou C, Parra J, Froment C, Rengel D, Burlet-Schiltz O, Marcoux J, Nigou J, Rivière M, Fabre E.(2023) Development of a novel target-based cell assay, reporter of the activity of mycobacterium tuberculosis protein-O-mannosyltransferase. Glycobiology 33, 1139 Pubmed
Tonini, L., Sadet, B., Stella, A., Bouyssie, D., Nigou, J., Burlet-Schiltz, O., Riviere, M. (2020) Potential Plasticity of the Mannoprotein Repertoire Associated to Mycobacterium tuberculosis Virulence Unveiled by Mass Spectrometry-Based Glycoproteomics. Molecules 25, 2348 (View)
Vetizou, M., Pitt, J.M., Daillere, R., Lepage, P., Waldschmitt, N., Flament, C., Rusakiewicz, S., Routy, B., Roberti, M.P., Duong, C.P., Poirier-Colame, V., Roux, A., Becharef, S., Formenti, S., Golden, E., Cording, S., Eberl, G., Schlitzer, A., Ginhoux, F., Mani, S., Yamazaki, T., Jacquelot, N., Enot, D.P., Berard, M., Nigou, J., Opolon, P., Eggermont, A., Woerther, P.L., Chachaty, E., Chaput, N., Robert, C., Mateus, C., Kroemer, G., Raoult, D., Boneca, I.G., Carbonnel, F., Chamaillard, M., Zitvogel, L. (2015) Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079-1084 (View)
Larrouy-Maumus, G., Skovierova, H., Dhouib, R., Angala, S.K., Zuberogoitia, S., Pham, H., Villela, A.D., Mikusova, K., Noguera, A., Gilleron, M., Valentinova, L., Kordulakova, J., Brennan, P.J., Puzo, G., Nigou, J., Jackson, M. (2012) A small multidrug resistance-like transporter involved in the arabinosylation of arabinogalactan and lipoarabinomannan in mycobacteria. J Biol Chem 287, 39933-39941 (View)
Reviews
Klionsky, D., …, Vergne, I., et al. (2021). Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition). Autophagy 17:1-382 (View)
Layre, E. (2020) Trafficking of Mycobacterium tuberculosis Envelope Components and Release Within Extracellular Vesicles: Host-Pathogen Interactions Beyond the Wall. Front Immunol 11, 1230 (View)
Bah, A., Vergne, I. (2017) Macrophage Autophagy and Bacterial Infections. Front Immunol 8, 1483 Pubmed (View)
Vergne, I., Lafont, F., Espert, L., Esclatine, A., Biard-Piechaczyk, M. (2017) Autophagy, ATG proteins and infectious diseases. Med Sci (Paris) 33, 312-318 Special Issue on “Autophagie”, editorial: Patrice Codogno et Guido Kroemer. (View)
Vergne, I., Gilleron, M., Nigou, J. (2014) Manipulation of the endocytic pathway and phagocyte functions by Mycobacterium tuberculosis lipoarabinomannan. Front Cell Infect Microbiol 4, 187 (View)
Layre, E., Mazurek, J., Gilleron, M. (2015) Glycolipid Presentation by CD1. In: eLS. John Wiley & Sons, Ltd: Chichester (View)
Ray, A., Cot, M., Puzo, G., Gilleron, M., Nigou, J. (2013) Bacterial cell wall macroamphiphiles: pathogen-/microbe-associated molecular patterns detected by mammalian innate immune system. Biochimie 95, 33-42 (View)
Publications in the past 5 years
2025
Géraud N., Rivière C., Falcou C., Cioci G., Froment C., Gervais V., Marcoux J., Gilleron M., Nigou J., Fabre E., Rivière M. (2025) Structural insights into the protein mannosyltransferase from Mycobacterium tuberculosis reveal a WW-domain-like protein motif in bacteria. Commun Biol 8, 1175 Pubmed
2024
Pouzol S, Khaja Mafij Uddin M, Islam A, Jabin MS , Nigou J, Banu S, Hoffmann J. (2024) Combination of exhaled breath condensate samples with lipoarabinomannan point of care assay for pulmonary tuberculosis (TB) diagnosis: protocol for a diagnostic accuracy study. BMJ Open. 14:e087026. doi: 10.1136/bmjopen-2024-087026
Kitzmiller CE, Cheng TY, Prandi J, Sparks IL, Moody DB, Morita YS. (2024) Detergent-induced quantitatively limited formation of diacyl phosphatidylinositol dimannoside in Mycobacterium smegmatis. J Lipid Res. Mar 22:100533. doi: 10.1016/j.jlr.2024.100533
Palčeková Z, De K, Angala SK, Gilleron M, Zuberogoitia S, Gouxette L, Soto-Ojeda M, Gonzalez-Juarrero M, Obregón-Henao A, Nigou J, Wheat WH, Jackson M. (2024) Impact of Methylthioxylose Substituents on the Biological Activities of Lipomannan and Lipoarabinomannan in Mycobacterium tuberculosis. ACS Infect Dis. Mar 21. doi: 10.1021/acsinfecdis.4c00079.
Lin Z, Kaniraj JP, Holzheimer M, Nigou J, Gilleron M, Hekelaar J, Minnaard AJ. (2024) Asymmetric Total Synthesis and Structural Revision of DAT2, an Antigenic Glycolipid from Mycobacterium tuberculosis. Angew Chem Int Ed Engl. Mar 8:e202318582. doi: 10.1002/anie.202318582
2023
Eckhardt E, Schinköthe J, Gischke M, Sehl-Ewert J, Corleis B, Dorhoi A, Teifke J, Albrecht D, Geluk A, Gilleron M, Bastian M. (2023) Phosphatidylinositolmannoside vaccination induces lipid-specific Th1-responses and partially protects guinea pigs from Mycobacterium tuberculosis challenge. Sci Rep. 13:18613.
Géraud N, Falcou C, Parra J, Froment C, Rengel D, Burlet-Schiltz O, Marcoux J, Nigou J, Rivière M, Fabre E.(2023) Development of a novel target-based cell assay, reporter of the activity of mycobacterium tuberculosis protein-O-mannosyltransferase. Glycobiology 33, 1139.
Palčeková Z, Obregón-Henao A, De K, Walz A, Lam H, Philp J, Angala SK, Patterson J, Pearce C, Zuberogoitia S, Avanzi C, Nigou J, McNeil M, Muñoz Gutiérrez JF, Gilleron M, Wheat WH, Gonzalez-Juarrero M, Jackson M. (2023) Role of succinyl substituents in the mannose-capping of lipoarabinomannan and control of inflammation in Mycobacterium tuberculosis infection. PLoS Pathog. 19:e1011636.
Nabeemeeah F, Sabet R, Moloantoa T, Waja Z, Pretorius Z, Majoro K, Letutu-Xaba M, Vilaplana C, Nigou J, Martinson N. (2023) Exhaled breath specimens subjected to point-of-care lipoarabinomannan testing. Int J Tuberc Lung Dis. 27:703-705.
Viljoen, A., Vercellone, A., Chimen, M., Gaibelet G., Mazères, S, Nigou, J., Dufrêne, Y.F. (2023) Nanoscale clustering of mycobacterial ligands and DC-SIGN host receptors are key determinants for pathogen recognition. Sci Adv 9, eadf9498
Aceves-Sánchez MJ, Barrios-Payán JA, Segura-Cerda CA, Flores-Valdez MA, Mata-Espinosa D, Pedroza-Roldán C, Yadav R, Saini DK, de la Cruz MA, Ares MA, Bielefeldt-Ohmann H, Baay-Guzmán G, Vergne I, Velázquez-Fernández JB, Barba León J, Hernández-Pando R. (2023) BCG∆BCG1419c and BCG differ in induction of autophagy, c-di-GMP content, proteome, and progression of lung pathology in Mycobacterium tuberculosis HN878-infected male BALB/c mice. Vaccine. May 8: S0264-410X(23)00489-9
Brown CM, Corey RA, Grélard A, Gao Y, Choi YK, Luna E, Gilleron M, Destainville N, Nigou J, Loquet A, Fullam E, Im W, Stansfeld PJ, Chavent M. (2023) Supramolecular organization and dynamics of mannosylated phosphatidylinositol lipids in the mycobacterial plasma membrane. Proc Natl Acad Sci U S A. 120 (5):e2212755120.
Abeliovich H, Debnath J, Ding WX, Jackson WT, Kim DH, Klionsky DJ, Ktistakis N, Margeta M, Münz C, Petersen M, Sadoshima J, Vergne I. (2023) Editorial: Where is the field of autophagy research heading? Autophagy. Jan 23;1-6
2022
Mosquera-Restrepo SF, Zuberogoïtia S, Gouxette L, Layre E, Gilleron M, Stella A, Rengel D, Burlet-Schilt O, Caro AC, Garcia LF, Segura C, Peláez Jaramillo CA, Rojas M, Nigou J. (2022) A Mycobacterium tuberculosis fingerprint in human breath allows tuberculosis detection. Nat Commun. 13(1):7751.
Correia-Neves M, Nigou J, Mousavian Z, Sundling C, Källenius G. (2022) Immunological hyporesponsiveness in tuberculosis: the role of mycobacterial glycolipids. Front Immunol. 13:1035122.
Daher W, Leclercq LD, Johansen MD, Hamela C, Karam J, Trivelli X, Nigou J, Guérardel Y, Kremer L. (2022) Glycopeptidolipid glycosylation controls surface properties and pathogenicity in Mycobacterium abscessus. Cell Chem Biol. 22:00125.
Viljoen A, Dufrêne YF, Nigou J. (2022) Mycobacterial Adhesion:From Hydrophobic to Receptor-Ligand Interactions. Microorganisms. 10:454.
James CA, Xu Y, Aguilar MS, Jing L, Layton ED, Gilleron M, Minnaard AJ, Scriba TJ, Day CL, Warren EH, Koelle DM, Seshadri C. (2022) CD4 and CD8 co-receptors modulate functional avidity of CD1b-restricted T cells. Nat Commun. 13:78.
2021
Sousa Silva C, Sundling C, Folkesson E, Fröberg G, Nóbrega C, Canto-Gomes J, Chambers B, Tadepally L, Brodin P, Bruchfeld J, Nigou J, Correia-Neves M & Källenius G. (2021) High dimensional immune profiling reveals different response patterns in active and latent TB following stimulation with mycobacterial glycolipids. Front Immunol. 12:727300.
Payros D, Alonso H, Malaga W, Volle A, Mazères S, Déjean S, Valière S, Moreau F, Balor S, Stella A, Combes-Soia L, Burlet-Schiltz O, Bouchez O, Nigou J, Astarie-Dequeker C, Guilhot C. (2021) Rv0180c contributes to Mycobacterium tuberculosis cell shape and to infectivity in mice and macrophages. PLoS Pathog. 17:e1010020.
Anso I, Basso LGM, Wang L, Marina A, Páez-Pérez ED, Jäger C, Gavotto F, Tersa M, Perrone S, Contreras FX, Prandi J, Gilleron M, Linster CL, Corzana F, Lowary TL, Trastoy B, Guerin ME. (2021) Molecular ruler mechanism and interfacial catalysis of the integral membrane acyltransferase PatA. Sci Adv. 7:eabj4565.
Jagannath C, McBride JW, Vergne I (2021) Editorial: The Autophagy Pathway: Bacterial Pathogen Immunity and Evasion. Front Immunol. 12:768935.
Layre E (2021) Targeted Lipidomics of Mycobacterial Lipids and Glycolipids. Methods Mol Biol. 2314:549.
Jøntvedt Jørgensen M, Grotle Nore K, Aass HC, Layre E, Nigou J, Mortensen R, Tasken K, Kvale D, Jenum S, Tonby K, Dyrhol-Riise AM (2021) Plasma LOX-products and monocyte signaling is reduced by adjunctive cyclooxygenase-2 inhibitor in a phase I clinical trial of tuberculosis patients. Front Cell Infect Microbiol 11:669623.
Lanéelle MA, Spina L, Nigou J, Lemassu A & Daffé M (2021) Lipid and lipoarabinomannan purification and characterization. Methods Mol Biol 2314:109.
Klionsky D, …, Vergne I, et al. (2021). Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition). Autophagy 17:1-382.
Felix J, Siebert C, Novion Ducassou J, Nigou J, Garcia PS, Fraudeau A, Huard K, Mas C, Brochier-Armanet C, Couté Y, Gutsche I & and Renesto P (2021) Structural and Functional Analysis of the Francisella Lysine Decarboxylase as a Key Actor in Oxidative Stress Resistance. Sci Rep 11:972.
2020
Marín Franco JL, Genoula M, Corral D, Duette G, Ferreyra M, Maio M, Dolotowicz MB, Aparicio-Trejo OE, Patiño-Martínez E, Charton A, Métais A, Fuentes F, Soldan V, Moraña EJ, Palmero D, Ostrowski M, Schierloh P, Sánchez-Torres C, Hernández-Pando R, Pedraza-Chaverri J, Rombouts Y, Hudrisier D, Layre E, Vérollet C, Maridonneau-Parini I, Neyrolles O, Sasiain MDC, Lugo-Villarino G, Balboa L (2020) Host-Derived Lipids from Tuberculous Pleurisy Impair Macrophage Microbicidal-Associated Metabolic Activity. Cell Rep 33:108547.
Källenius G, Nigou J, Cooper A & Correia-Neves M (2020) Editorial: Mycobacterial Glycolipids-Role in Immunomodulation and Targets for Vaccine Development. Front Immunol 11:603900.
Camacho F, Moreno E, Garcia-Alles LF, Santiago GC, Gilleron M, Vasquez A, Choong YS, Reyes F, Norazmi N, Sarmiento ME & Acosta A (2020) A Direct Role for the CD1b Endogenous Spacer in the Recognition of a Mycobacterium tuberculosis Antigen by T-Cell Receptors. Front Immunol 11:566710.
Layre E. (2020) Trafficking of Mycobacterium tuberculosis Envelope Components and Release Within Extracellular Vesicles: Host-Pathogen Interactions Beyond the Wall. Front Immunol 11:1230.
Palčeková Z, Gilleron M, Angala SK, Belardinelli JM, McNeil M, Bermudez LE, Jackson M. (2020) Polysaccharide Succinylation Enhances the Intracellular Survival of Mycobacterium abscessus. ACS Infect Dis 6:2235-2248.
Le Moigne V, Roux A-L, Jobart-Malfait A, Blanc L, Chaoui K, Burlet-Schiltz O, Gaillard J-L, Canaan S, Nigou J and Herrmann J-L. (2020) A TLR2-Activating Fraction From Mycobacterium abscessus Rough Variant Demonstrates Vaccine and Diagnostic Potential. Front Cell Infect Microbiol 10:432.
Tonini L, Sadet B, Stella A, Bouyssié D, Nigou J, Burlet-Schiltz O & Rivière M. (2020) Potential Plasticity of the Mannoprotein Repertoire Associated to Mycobacterium tuberculosis Virulence Unveiled by Mass Spectrometry-Based Glycoproteomics. Molecules 25:2348.
Le Moigne V , Raynaud C, Moreau F, Dupont C, Nigou J, Neyrolles O, Kremer L & Herrmann J-L. (2020) Efficacy of Bedaquiline, Alone or in Combination With Imipenem, Against Mycobacterium Abscessus in C3HeB/FeJ Mice. Antimicrob Agents Chemother 64:e00114-20.
Dubé J-Y , McIntosh F, Zarruk JG, David S, Nigou J, & Behr MA. (2020) Synthetic Mycobacterial Molecular Patterns Partially Complete Freund’s Adjuvant. Sci Rep 10:5874.
Bah A, Sanicas M, Nigou J, Guilhot C, Astarie-Dequeker C, & Vergne I. (2020) Lipid virulence factors of Mycobacterium tuberculosis exert a multilayered control of autophagy-related pathways in infected human macrophages. Cells 9:666.
2019
Galais M, Pradel B, Vergne I, Robert-Hebmann V, Espert L, Biard-Piechaczyk M. (2019) LAP (LC3-associated phagocytosis): phagocytosis or autophagy? Médecine/Sciences 35:635-642.
Palčeková Z, Angala SK, Belardinelli JM, Eskandarian HA, Joe M, Brunton R, Rithner C, Jones V, Nigou J, Lowary TL, Gilleron M, McNeil M, Jackson M (2019) Disruption of the SucT acyltransferase in Mycobacterium smegmatis abrogates succinylation of cell envelope polysaccharides. J Biol Chem 294:10325-35.
Souriant S, Balboa L, Dupont M, Pingris K, Kviatcovsky D, Cougoule C, Lastrucci C, Bah A, Gasser R, Poincloux R, Raynaud-Messina B, Al Saati T, Inwentarz S, Poggi S, Moran EJ, Gonzalez-Montaner P, Corti M, Lagane B, Vergne I, Allers C, Kaushal D, Kuroda MJ, del Carmen Sasiain M, Neyrolles O, Maridonneau-Parini I, Lugo-Villarino G, Vérollet C (2019) Tuberculosis exacerbates HIV-1 infection through IL-10/STAT3-dependent tunneling nanotube formation in macrophages. Cell Rep 26:P3586.
Shahine A, Reinink P, Reijneveld JF, Gras S, Holzheimer M, Cheng TY, Minnaard AJ, Altman JD, Lenz S, Prandi J, Kubler-Kielb J, Moody DB, Rossjohn J & Van Rhijn I (2019) A T-cell receptor escape channel allows broad T-cell response to CD1b and membrane phospholipids. Nat Commun 10:56.
Law C.T., Camacho F., Garcia-Alles L.F., Gilleron M., Sarmientol M., Norazmi M.N., Acosta A., Choong Y.S. (2019) Interactions of domain antibody (dAbk11) with Mycobacterium tuberculosis Ac2SGL in complex with CD1b. Tuberculosis 114:9-16.
Patents
Dendrimères à terminaison saccharide à visée anti-inflammatoire. Blattes, E., Puzo, G., Prandi, J., Nigou, J., Vercellone, A., Majoral, J.P., Turrin, C.O., Caminade A.M. Patent n° 10 56902 (FR, 31/08/2010), 11764827.9 (EU, 31/08/2011), 13/819872 (USA, 31/08/2011).
Pharmaceutical compositions comprising actinomycete glycerol acyl derivatives antigens, their process of extraction, and their use against tuberculosis. Puzo, G., Layre, E., Gilleron, M., Prandi, J., Stenger, S., De Libero, G. EP20070291249, deposited 12/10/2007, extended 13/10/2008.
Composition pour la prévention et/ou le traitement des maladies associées à la surexpression du TNF et/ou de l’IL12. Quesniaux, V., Gilleron, M., Puzo, G., Nigou, J. FR20060010136, deposited 20/11/2006, extended 20/11/2007.
Sulfoglycolipid antigens, their process of preparation, and their use against tuberculosis. Puzo, G., Prandi, J., Gilleron, M., De Libero, G., Guiard, J., Mori, L., Paoletti, S. EP20070290097, deposited 24/01/2007, extended 13/10/2008.
Sulfoglycolipid antigens, their extraction from Mycobacterium tuberculosis, and their use against tuberculosis. Puzo, G., Gilleron, M., Stenger, S., De Libero, G. EP20030290965, deposited 18/04/2003.
PhDs
- Nicolas Géraud (PhD 2021): Engineer, INSERM, Toulouse, France
- Pierre Boyer (PhD 2021): CBU Granits, Villefranche de Rouergue, France
- Christophe Carrat (PhD 2019): (1) Quality Engineer, Thermo Fisher Scientific, Nîmes, France; (2) Quality Engineer, Sanofi, Caumont-sur-Durance, France, since 2022
- Camille Robert (PhD 2017): (1) Research scientist, Evotec, Manchester, UK; (2) SNIPR Biome, Copenhagen, Denmark, since 2020
- Aicha Bah (PhD 2017): Medical science liaison, Terumo BCT, Brussels, Belgium
- Jin Wang (PhD 2015): Associate Professor Yancheng University, China
- Alexiane Decout (PhD 2015): (1) Research scientist, GLYcoDiag, Orléans, France; (2) Post-doctoral fellow Université Polytechnique Fédérale de Lausanne, Switzerland, 2017-20; (3) Research Associate, Imperial College, London, UK, since 2020
- Landry Blanc (PhD 2014): (1) Post-doctoral fellow Rutgers University, Newark, USA; (2) Post-doctoral fellow CNRS-Université de Bordeaux, France, since 2019; (3) CNRS research associate, IPBS, Toulouse, France, since 2023
- Laure Tonini (PhD 2014): (1) Engineer, CNRS-INSERM-Université de Toulouse; (2) Innovation finance consultant Leyton, Toulouse, France, since 2018
- Romain Castanier (PhD 2013): Quality control manager, dairy facility, Aveyron, France
- Diane Cala-De Paepe (PhD 2011): Engineer, CNRS-Université de Lyon, France
- Emilyne Blattes (PhD 2010): (1) Post-doctoral fellow University of Bern, Switzerland; (2) Medical science liaison, Innovative Medecine for Tuberculosis, Lausanne, Switzerland, 2016-19; (3) R&D Manager, 2 Bridge CVBA, Paris, France, since 2020
- Aurélie Ray (PhD 2010): (1) Post-doctoral fellow Trudeau Institute, Saranac Lake, USA; (2) Research scientist, Transgène, Lyon, France, 2013-2019; (3) Research scientist, Evotec, Lyon, France, since 2020
- Gérald Larrouy-Maumus (PhD 2009): (1) Post-doctoral fellow National Institute for Medical Research, MRC, London, UK; (2) Group leader Imperial College London, UK, since 2014
- Chia-Fang Liu (PhD 2008): Miidex, Toulouse, France
- Emilie Layre (PhD 2008): (1) Post-doctoral fellow Harvard Medical School, Boston, USA; (2) CNRS research associate, IPBS, Toulouse, France, since 2014
Post-docs
- Albertus Viljoen 2024: CNRS Research associate, IPBS, Toulouse, France, since 2024
- Deisy Rocha Silva 2022 : Maternity leave
- Jolanta Mazurek 2017: (1) Scientist Selvita, Cracow, Poland; (2) Team leader, Ryvu Therapeutics, Cracow, Poland, since 2018
- Naila Mebarek 2016: Assistant Professor Université de Bordeaux, France
- Sandro Silva-Gomes 2016: Research scientist, GSK, Stevenage Herts, UK
- Bachir Saadeh 2015: Sales engineer 10x Genomics, Toulouse, France
- Shiv Dubey 2009: Assistant Professor Govind Ballabh Pant University, Uttarakhand, India
- Marlène Cot 2009: Research scientist CRITT Bio industries, Toulouse, France
- Sandrine Bouhet 2008: Head scientific program coordinator, Europa Group, Toulouse, France
- Eliette Darthuy 2008: AQSE, Toulouse, France
Research assistants
- Hanamée Faugeras 2025: PhD, see2cure, Université de Toulouse
- Florian Boulée 2024 : Engineer in molecular biology and HTS, CNRS-Aix-Marseille Université, Marseille, France
- Sébastien Nicolas 2024 : Medical science liaison professional training
- Lucie Gouxette 2022: Engineer cell biology, Evotec, Toulouse, France
- Annie Behar 2021: (1) Engineer in animal experimentation, VibioSphen, Toulouse, France; (2) Engineer in animal experimentation, Evotec, Toulouse, France, since 2021
- Camille Falcou 2020: Engineer in molecular biology, CNRS-Aix-Marseille Université, Marseille, France
- Fabien Riols 2018: Engineer in metabolomics, University of Munich, Germany
- Florence Iehl 2017: Engineer in biochemistry, Evotec, Toulouse, France
- Laure Ducassé 2015: Engineer in animal experimentation InvivoGen, Toulouse, France
- Bérengère Ferrié 2015: Technician chemist Evotec, Toulouse, France
- Marion Bonhomme 2015: Technician chemist Adocia, Lyon, France
- Anne-Sophie Bardou 2015: Social supervisor Midi-Pyrénées Expertise, Toulouse, France
- Sophie Zuberogoitia 2015: Engineer biochemist, Enobraq, Toulouse, France; 2019: CNRS Engineer, IPBS, Toulouse, France
- Marie Gonzales 2015: Technician chemist Evotec, Toulouse, France
- Delphine Bordignon 2014: Technician chemist Solvionic, Toulouse, France
- Cécile Glénat 2013: Technician chemist Etienne Lacroix Group, Sainte-Foy-de-Peyrolières, France
- Benjamin Gau 2010: Quality Specialist SGS, Montréal, Canada

